_Overview.jpg.webp)
Telomeric repeat–containing RNA (TERRA) is a long non-coding RNA transcribed from telomeres - repetitive nucleotide regions found on the ends of chromosomes that function to protect DNA from deterioration or fusion with neighboring chromosomes. TERRA has been shown to be ubiquitously expressed in almost all cell types containing linear chromosomes - including humans, mice, and yeasts. While the exact function of TERRA is still an active area of research, it is generally believed to play a role in regulating telomerase activity as well as maintaining the heterochromatic state at the ends of chromosomes. TERRA interaction with other associated telomeric proteins has also been shown to help regulate telomere integrity in a length-dependent manner.
Due to the breadth of roles in which TERRA is implicated for maintaining the genomic integrity at the ends of chromosomes, TERRA dysfunction has also been shown to be associated with a number of disease states, including a number of syndromes related to inappropriate telomere shortening and cellular aging, and the progression of cancer. As such, pharmacologic compounds modulating TERRA transcription as well as its other direct/indirect interactions may be a viable method for the development of new therapeutic strategies.
Structure & Biogenesis
Transcription From DNA
TERRA is an evolutionarily conserved long-non-coding RNA found in many nucleus-containing eukaryotic cells such as chicken (Gallus gallus domesticus),[1] humans (Homo sapiens), budding yeast (Schizosaccharomyces cerevisiae), fission yeast (Schizosaccharomyces pombe), mice (Mus musculus), zebrafish (Danio rerio), and various plants (Arabidopsis thaliana, et cetera).[2][3][4][5] TERRA is transcribed from DNA as a continuous sequence in most tissues from the subtelomeric and telomeric regions at the end of linear chromosomes. Transcription occurs in the 5' to 3' direction (from the centromere to telomere) and TERRA transcripts are heterogeneous in length, ranging anywhere from 100-bases up to 9-kilobases in length for human cells.[6] The majority of TERRA transcription has been shown to be carried out by RNA Polymerase II (RNAPII),[3][4] although RNA Polymerase I and III may also have functions in TERRA transcription for mammals.[7]
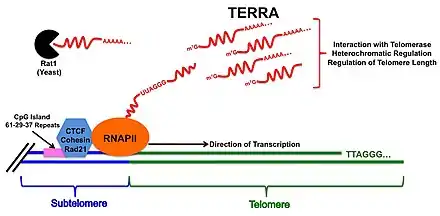
While the exact transcription start site is still a matter of debate, it is widely held that TERRA transcription begins in the subtelomeric region at the end of chromosomes, just before the tandem repeats of the telomere begin. In humans, at least 20 of 23 chromosome ends have been shown to contain CpG islands, which are generally found at or near transcription start sites in the subtelomeric region. Within these CpG island are three repetitive DNA tracts occurring in sequence: a centromere-proximal tract of repeated 61-base-pair (bp) units; second, a more distal tract of 29-bp units; and a third 37-bp unit tract. These highly conserved repetitive elements are known as 61-29-37 Repeats. Additionally, the chromatin organization factors CTCF (CCCTC-Binding Factor), Cohesin, and Rad21 - which are known to aid in the recruitment of RNAPII[8] - have also been shown to bind within 1-2 kilobases of the telomeric tracks just upstream of these CpG islands.[9] While not definitely known, the proximity of these factors immediately upstream of the proposed TERRA transcription start site suggest that they may function as promoter elements for the start of TERRA transcription.[6]
Heterogeneity of Transcript Length
The wide range in length of TERRA transcripts is believed to primarily stem from either different sites of transcription termination within the telomere tract or variation in 3'-end processing. As telomeres do not contain a canonical stop codon signaling the end of RNA transcription, it remains unclear exactly where transcription ceases along the telomere tracks. Similarly to most mRNAs transcribed in the nucleus, nearly all TERRA transcripts contain a 7-methylguanosine (m7G) cap structure at their 5' end.[10] Therefore, the vast majority of post-transcriptional variation occurs at the 3' end of the transcript; however this has been shown to vary between species, as well as within individual cells. In humans, approximately 7 to 10% of TERRA transcripts have been shown to be post-transcriptionally polyadenylated,[11][3] whereas almost all TERRA transcribed in yeast contains a poly(A) tail[4] added by the polyadenylation polymerases Pap1 (in S. cerevisiae)[4] or Cid12/Cid14 (in S. pombe).[12] Both of these post-transcriptional modifications contribute to stabilizing TERRA within the cell (half-life ~8 hours) compared to non-modified transcripts (half-life ~3 hours).[3][10] Additionally, the presence of a polyadenylated tail also appears to act as a determinant of TERRA localization within the nucleus. TERRA transcripts containing a poly(A) tail are generally found "free" within the nucleoplasm while transcripts lacking a polyadenylated tail are divided between the nucleoplasm (60%) and a chromatin-bound state (40%).[10]
Factors Regulating Transcription
A number of proteins shown to associate with the ends of telomeres are involved in modulating TERRA transcription levels within cells. Factors involved in modulating the heterochromatic state at telomeres, such as histone deacetylase,[11] SUV39H1 H3K9 histone methyltransferase,[3] and DNA methyltransferase 3b (DNMT3B) in human cells,[13] play an important role in regulating total TERRA expression. Additionally, the Shelterin component TRF1 has been shown to support TERRA transcription.[3] As part of the six-protein complex involved with protecting the ends of telomeres, TRF1 has been shown to promote progression of TERRA's primary transcription polymerase RNAPII through telomere tracts, although it has not been shown to promote direct recruitment of the polymerase.
In yeast cells, TERRA stability has been shown to be tightly regulated within the nucleus by the 5' to 3' exonuclease Rat1 (human ortholog Xrn2) through interaction with the proteins Rap1 and Rif1/2.[4][14] It remains unclear whether TERRA transcripts remain tethered to the telomeres from which they are transcribed, acting in cis, or whether they may move from one telomere to another by some unknown mechanism to regulate their functions (acting in trans).
Other Telomeric RNA Transcripts
While TERRA currently represents the primary subject of study for RNA transcripts originating from telomeres, a wide variety of alternative transcripts have also been identified as part of the yeast telomeric transcriptome: a transcript complementary to the 5' subtelomeric sequence known as ARRET; a subtelomeric transcript complementary to ARRET known as αARRET; and a C-rich telomeric repeat-containing transcript known as ARIA.[12][15]
Functions of TERRA
Interaction with Telomerase
Telomerase is the ribonucleoprotein responsible for adding species-dependent tandem repeat sequences (TTAGGG in humans) to the ends of telomeres. These telomeric repeats function to protect the ends of the chromosome from DNA damage or end-to-end fusion with adjacent chromosomes. Since evidence has also shown that TERRA expression is regulated in a telomere-length-dependent manner,[3] it is believed that TERRA may play an important role along with telomerase to modulate the length of telomere repeats on chromosome ends. Generally speaking, cells with long telomeres exhibit greater TERRA expression while short telomeres have relatively lower TERRA expression. At the same time, telomerase activity is known to be at its greatest when telomeres are short and at its lowest activity when telomeres are long. It can therefore be proposed that TERRA acts as a negative regulator of telomerase activity in cells with long telomeres,[3][4][16] while functioning as a positive regulator of telomerase activity at short telomeres[8] - in conjunction with relative TERRA expression for each telomeric state.
TERRA & Telomerase at Long Telomeres
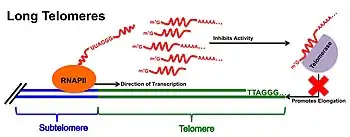
At cells with long telomeres, TERRA is proposed to act as a negative regulator of telomerase activity. This direct inhibitor function acts through base-pairing of the tandem repeats found throughout TERRA's 3'-end to the complementary RNA template region of telomerase.[6] The complementary base pairing of the TERRA transcript may work to compete with telomerase's DNA substrate when TERRA expression is at its highest, preventing telomerase from further lengthening telomeres. TERRA may also interact with the catalytic reverse transcriptase subunit (TERT) of telomerase, thereby acting as an allosteric inhibitor of telomerase function and preventing the addition of further telomeric repeats. This model of TERRA-telomerase interaction is supported by experimental evidence where TERRA-mimicking (UUAGGG)3 oligonucleotides have been shown to lead to complete inhibition of telomerase activity in vitro;[3][17] however evidence of this relationship in vivo remains elusive.
TERRA & Telomerase at Short Telomeres
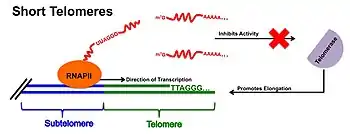
In seemingly contradictory fashion, it is believed that TERRA may instead coordinate the recruitment of telomerase and encourage subsequent telomere elongation for cells containing short telomeres. Specifically in short telomeres of yeast, evidence suggests that TERRA molecules help to form clusters with the yeast telomerase RNA TLC1. These clusters, known as T-Recs, have been shown to preferentially localize to short telomeres and may therefore encourage telomerase-based elongation.[18] Furthermore, proteins that preferentially bind the UUAGGG repeats within TERRA molecules, such as the heterogeneous nuclear ribonucleoprotein hnRNP1A, have been shown to prevent the direct TERRA-telomerase inhibition found at short telomeres, thus promoting telomerase-mediated elongation.
The true nature of TERRA's interaction with telomerase remains incompletely described. However, it remains clear that the ability of telomerase to add telomeric repeats to the ends of chromosomes is intricately intertwined with its interaction with TERRA.
Heterochromatin Regulation
In general, long non-coding RNAs are known to mediate epigenetic changes on chromatin by functioning as recruiters for chromatin-modifying enzymes to genomic loci. TERRA is thought to play a similar role, recruiting both heterochromatic proteins and associated chromatin-remodeling complexes to telomeres, helping to establish or maintain heterochromatin formation.
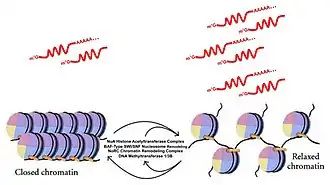
One manner in which chromatin maintains its condensed hetererochromatic state at the telomere is through the enzymes DNA methyltransferases 1 and 3b (DNMT1/3b). Experiments where these factors have been depleted result in increased TERRA expression levels,[19] suggesting that methylation status of the subtelomeric region may help to regulate the expression of TERRA. Further post-translational modifications of telomeric histones may also play a pivotal role in TERRA regulation. Decreased density of trimethylated histones H3K9me3 and H4K30me3 have been shown to correlate with increased TERRA levels.[20] RNA pulldown experiments have also shown that TERRA associates with heterochromatin proteins, including HP1α, HP1β, and the origin recognition complex (ORC).[21] These factors are known to play a role in establishment of heterochromatin and subsequent transcriptional silencing. TERRA-mimicking oligonucleotides have also been shown to associate with other chromatin remodeling complexes such as the NuA Histone Acetyltransferase Complex, BAF-Type SWI/SNF Nucleosome Remodeling Complex, and the NoRC Chromatin Remodeling Complex.[22] It is believed that TERRA may work to stabilize the interaction of these factors with chromatin to promote heterochromatin formation.
Overall, evidence has shown a complex relationship whereby TERRA can both regulate the heterochromatic state and be regulated by the heterochromatic state. Increased density of repressive histone marks that cause chromatin to become more condensed correlate with observed decreases in TERRA expression. Concurrently, epigenetic changes that result in chromatin becoming more euchromatic or "open" in nature correlates with increased TERRA expression. Listed above are factors shown to contribute to modulating TERRA expression, but this is by no means an exhaustive list.
Regulation of Telomere Length
TERRA has been shown to exist exclusively within the nucleus of eukaryotic cells where it specifically associates with the ends of chromosomes at the telomere. While TERRA has not been shown to be an essential or permanent component of telomere chromatin, it is believed that TERRA's primary function is to transiently maintain the structural integrity at telomeres, either through direct interaction with telomeric DNA or by the binding of associated telomeric proteins.
Direct Telomeric Interactions
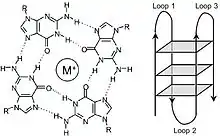
Data has shown that TERRA interacts with telomeric DNA to form G-Quadruplex structures both in vitro and in vivo.[23][24] These structures form due to the abundance of guanine found in the TERRA transcript and their ability to associate through Hoogsteen hydrogen bonding. The resulting square planar structure is known as a guanine tetrad (also known as a G-tetrad or G-quartet) and when two or more guanine tetrads stack on top of each other, they form a G-quadruplex. These complex structures have been shown help to modulate telomere length through inhibition telomerase's ability to add tandem TTAGGG repeats, specifically in cells with long telomeres. This inhibitive effect seems to function in a telomere length-dependent manner;[3] that is, TERRA levels inversely correlate with increased activity of telomere elongation. In general, TERRA has been shown to be most abundant in cells with long telomeres,[2][3] while cells with short telomeres express comparatively lower levels of transcript expression. There is also evidence that overexpression of TERRA in human cells can help promote telomere processing by inhibiting the 5'-3' exonuclease Exo1 through the Ku70/80 dimer.[22][25]
TERRA expression has also been shown to be cell cycle regulated via the chromatin-remodeling protein ATRX, with the highest expression of TERRA being present at the G1 phase following mitosis and before progressively declining to its lowest in S phase.[10] Therefore, TERRA expression may play an important role in the proper orchestration of DNA replication, as TERRA may hybridize with single-stranded DNA during replication.[21][26][27][28] In cells with long telomeres and high TERRA expression, this could potentially lead to replicative arrest and the eventual collapse of the replication fork, resulting in double-stranded breaks at the telomere.[26] This model is supported by evidence in other regions of the genome where persistent DNA-RNA hybrids lead to increased DNA damage.[29][30][31]

TERRA has also been indicated to be involved with the stabilization of the telomere-protecting Shelterin complex at the ends of telomeres. Evidence shows that TERRA functions together with hnRNPA1 to displace RPA from single-stranded DNA following replication and promotes the loading of the Shelterin component POT1 onto telomere ends.[32] This helps to stabilize the Shelterin complex so that it can properly cap the ends of chromosomes and protect them from recognition by DNA damage signaling pathways.
TERRA and the Alternative Lengthening of Telomeres (ALT) Pathway
In cells that lack telomerase expression, it is well-established that most cells instead maintain the ends of their telomeres through the recombination-based Alternative Lengthening of Telomeres (ALT) pathway. It is proposed that TERRA may work to delay the onset of cellular senescence in these cells and instead promote mechanisms of homologous recombination, thereby driving the ALT phenotype as an alternative means of maintaining telomere ends.[33] In support of this model, TERRA expression has also been shown relate to cellular ALT-status. Cells that have been experimentally confirmed to utilize the ALT pathway have markedly higher levels of TERRA expression compared to non-ALT cells.[33] The specific nature of the complex interactions involved in this proposed model remain an area of active research.
Indirect Telomeric Interactions
TERRA has further been shown to play a role in a number of interactions with proteins functioning at the ends of telomeres. Association with nonsense-mediated RNA decay (NMD) machinery such as UPF1, SMG1 and EST1A/SMG6[2][34] and members of the heterogeneous nuclear ribonucleoprotein (hnRNP) family have also been shown to regulate the displacement of TERRA from the ends of telomeres. The ability of these mechanisms to actively coordinate the displacement of TERRA provides evidence that TERRA is not a stable constituent of the telomeric chromatin, but not necessarily unstable itself.[2][35] Instead, the regulated control of TERRA displacement suggests a link between TERRA localization at the telomere and overall genome integrity. For example, in short telomeres with low TERRA expression, UPF1 and other SMG factors may work in concordance with DNA Polymerase δ to remove TERRA from the telomere during replication and alleviate fork stalling to allow for progression of replicative machinery. Additionally, replication fork stalling and SMG factor recruitment may also aid in telomerase localization to the telomeres where de novo repeats may be added to promote elongation of short chromosomes.
Proposed Role in Human Disease
While there are no diseases directly attributed to TERRA dysfunction, a number of diseases related to inappropriate telomere shortening and cellular aging may have dysfunctional TERRA expression as one of their causative factors. Such examples include cancer development and progression,[36][37] Dyskeratosis Congenita (DC),[38] Idiopathic Pulmonary Fibrosis (IPF),[39] Aplastic Anemia (AA),[40] Astrocytoma,[41] and Cryptogenic Liver Cirrhosis.[42]
Immunodeficiency, Centromeric Region Instability, Facial Anomalies (ICF Syndrome)
ICF is a rare autosomal recessive disease arising from deficiencies or mutations in DNA Methyltransferase 3b (DNMT3b), which acts as a key modulator in maintaining heterochromatin at telomeres.[43] As a result, many ICF patients experience spontaneous genomic instability in various tissues and systemic immunodeficiency. Contrary to traditionally low TERRA levels at short telomeres it has recently been shown that ICF patients display increased TERRA transcription[13] at critically shortened telomeres. However, this is likely the result DNMT3b deficiencies or mutations causing hypomethylation of the CpG islands within the subtelomeric region, allowing for increased TERRA transcription.[13][44] Further evidence also shows a decrease in trimethylated histone H3K9 at the telomeres and loss of binding of the Shelerin-component TRF2 at the subtelomeres.[44] As a result, the increased TERRA expression further contributes to abnormal shorting of the telomere, leading to the spontaneous genomic instability and systemic immunodeficiency. At the minimum, there appears to be a causal link between DNA methylation, TERRA regulation, and telomere length - although the relationship is currently not well understood.
In Cancer
One of the most distinct differences in TERRA expression exists between cancer cells, which exhibit significantly increased TERRA levels, and normal somatic tissues. This observation suggests that high levels of TERRA expression must evolutionarily benefit cancer progression in some way, possibly in their ability to promote unchecked cellular proliferation and induce cellular transformation. Even within subtypes of cancer cells, there exist differences in TERRA expression based on the mechanism by which cells maintain the length of their telomeres; TERRA levels have been shown to be consistently lower in telomerase positive cells compared to cells that utilize the homologous recombination-based Alternative Lengthening of Telomeres (ALT) mechanism.[33][41] TERRA may give ALT-positive cancer cells a means to promote genomic instability at telomeres and allow for this recombination-based method of elongation. It remains unclear in this model if TERRA is a cause or consequence of the ALT pathway. Methylation status of the subtelomeric histones may also play an important role in TERRA expression in cancer cells. Many cancer cell subtypes are known to be heavily methylated at the CpG islands of the subtelomeric promoter region, resulting in increased transcription of TERRA.[45]
As a Target for Small Molecules
There are currently no known inhibitors that specifically target TERRA in cells. However, relating to many of the diseases and syndromes listed above, it is believed that targeted transcriptional silencing of TERRA may potentially provide a viable mechanism for treatment of cells with high levels of TERRA expression.
Increasing the activity of TERRA-repressive histone deacetylases such as SIRT1 and SIRT6 through the use of allosteric activators may be prime candidates for the active repression of TERRA transcription.[14] Commercially available sirtuin-activating compounds (STACs) such as Resveratrol, Butein, Piceatannol, Isoliquiritigenin, Fisetin, and Quercetin are examples of small molecules that may target this axis.[46] Targeting TERRA with antisense RNA or RNAi technology may also be a way to promote proper telomere function and preserve telomere length. However it is important to note that complete abolishment of TERRA transcription must be avoided, as TERRA's association with the shelterin component POT1 plays a key role in recruitment of the complex to the ends of telomeres. As a further consideration, decreased TERRA expression on an organismal level may increase the replicative capacity of all actively dividing cells, thus potentially promoting tumorigenesis. Only a therapy targeted to cells with high levels of TERRA expression should be considered in these cases.
Since TERRA is also proposed to have a role in telomerase inhibition, factors increasing TERRA transcription may also be useful in select cases where aberrant telomerase activity is a contributing factor of disease. General HDAC inhibitors or specific sirtuin inhibitors such as Nicotinamide or Cambinol may prove to be viable treatment options in these cases. Trichostatin A (a HDAC1/2 inhibitor) has already been shown to significantly increase TERRA levels in human cancer cells.[11] It is important to note that TERRA upregulation alone has not been shown to be sufficient to induce telomere loss, so these drugs may need to be given in combination with other therapies, such as telomerase inhibitors.
See also
References
- ↑ Solovei, I.; Gaginskaya, E. R.; Macgregor, H. C. (November 1994). "The arrangement and transcription of telomere DNA sequences at the ends of lampbrush chromosomes of birds". Chromosome Research. 2 (6): 460–470. doi:10.1007/BF01552869. ISSN 0967-3849.
- 1 2 3 4 Azzalin CM, Reichenbach P, Khoriauli L, Giulotto E, Lingner J. Telomeric repeat containing RNA and RNA surveillance factors at mammalian chromosome ends. Science 2007, 318:798–801.
- 1 2 3 4 5 6 7 8 9 10 11 Schoeftner S, Blasco MA. Developmentally regulated transcription of mammalian telomeres by DNA- dependent RNA polymerase II. Nat Cell Biol 2008, 10:228–236.
- 1 2 3 4 5 6 Luke B, Panza A, Redon S, Iglesias N, Li Z, Lingner J. The Rat1p 5’ to 3’ exonuclease degrades telomeric repeat-containing RNA and promotes telomere elongation in Saccharomyces cerevisiae. Mol Cell 2008, 32:465–477.
- ↑ Vrbsky J, Akimcheva S, Watson JM, Turner TL, Daxinger L, Vyskot B, Aufsatz W, Riha K. siRNA–mediated methylation of Arabidopsis telomeres. PLoS Genet 2010, 6:e1000986.
- 1 2 3 André Maicher, Lisa Kastner & Brian Luke (2012) Telomeres and disease: Enter TERRA, RNA Biology, 9:6, 843-849, doi:10.4161/rna.20330.
- ↑ Déjardin, Jérôme & E Kingston, Robert. (2009). Purification of Proteins Associated with Specific Genomic Loci. Cell. 136. 175-86. 10.1016/j.cell.2008.11.045.
- 1 2 Cusanelli Emilio, Chartrand Pascal. Telomeric noncoding RNA : telomeric repeat‐containing RNA in telomere biology. WIREs RNA 2014, 5: 407-419. doi:10.1002/wrna.1220
- ↑ Deng Z, Wang Z, Stong N, Plasschaert R, Moczan A, Chen H-S, Hu S, Wikramasinghe P, Davuluri RV, Bartolomei MS, etal. A role for CTCF and cohesin in subtelomere chromatin organization, TERRA transcription, and telomere end protection. EMBO J 2012, 31:4165–4178.
- 1 2 3 4 Porro A, Feuerhahn S, Reichenbach P, Lingner J. Molecular dissection of telomeric repeat-containing RNA biogenesis unveils the presence of distinct and multiple regulatory pathways. Mol Cell Biol 2010, 30:4808–4817.
- 1 2 3 Claus M. Azzalin & Joachim Lingner (2008) Telomeres: The silence is broken, Cell Cycle, 7:9, 1161-1165, doi:10.4161/cc.7.9.5836
- 1 2 Bah A, Wischnewski H, Shchepachev V, Azzalin CM. The telomeric transcriptome of Schizosaccharomyces pombe. Nucleic Acids Res 2012, 40:2995–3005.
- 1 2 3 Yehezkel S, Segev Y, Viegas-Pe ́quignot E, Skorecki K, Selig S. Hypomethylation of subtelomeric regions in ICF syndrome is associated with abnormally short telomeres and enhanced transcription from telomeric regions. Hum Mol Genet 2008, 17:2776–2789.
- 1 2 N. Iglesias, et al., Subtelomeric repetitive elements determine TERRA regulation by Rap1/Rif and Rap1/Sir complexes in yeast, EMBO Rep. 12 (6) (2011) 587–593.
- ↑ Greenwood J, Cooper JP. Non-coding telomeric and subtelomeric transcripts are differentially regulated by telomeric and heterochromatin assembly factors in fission yeast. Nucleic Acids Res 2012, 40:2956–2963.
- ↑ Redon S, Reichenbach P, Lingner J. Protein RNA and protein protein interactions mediate association of human EST1A/SMG6 with telomerase. Nucleic Acids Res 2007; 35:7011-22; PMID 17940095; http:// dx.doi.org/10.1093/nar/gkm724.
- ↑ Redon S, Reichenbach P, Lingner J. The non-coding RNA TERRA is a natural ligand and direct inhibitor of human telomerase. Nucleic Acids Res 2010, 38:5797–5806.
- ↑ Cusanelli E, Romero CAP, Chartrand P. Telomeric noncoding RNA TERRA is induced by telomere shortening to nucleate telomerase molecules at short telomeres. Mol Cell 2013, 51:780–791.
- ↑ Farnung BO, Brun CM, Arora R, Lorenzi LE, Azzalin CM. Telomerase efficiently elongates highly transcribing telomeres in human cancer cells. PLoS ONE 2012, 7:e35714.
- ↑ Marion RM, Strati K, Li H, Tejera A, Schoeftner S, Ortega S, Serrano M, Blasco MA. Telomeres acquire embryonic stem cell characteristics in induced pluripotent stem cells. Cell Stem Cell 2009, 4:141–154.
- 1 2 Deng Z, Norseen J, Wiedmer A, Riethman H, Lieberman PM. TERRA RNA binding to TRF2 facilitates heterochromatin formation and ORC recruitment at telomeres. Mol Cell 2009, 35:403–413.
- 1 2 Scheibe M, Arnoult N, Kappei D, Buchholz F, Decottignies A, Butter F, Mann M. Quantitative interaction screen of telomeric repeat-containing RNA reveals novel TERRA regulators. Genome Res 2013.
- ↑ Yan Xu, Kuniyuki Kaminaga, and Makoto Komiyama. G-Quadruplex Formation by Human Telomeric Repeats-Containing RNA in Na+ Solution. Journal of the American Chemical Society. 2008 130 (33), 11179-11184. doi:10.1021/ja8031532
- ↑ Schaffitzel C, Berger I, Postberg J, Hanes J, Lipps HJ, Plückthun A. In vitro generated antibodies specific for telomeric guanine-quadruplex DNA react with Stylonychia lemnae macronuclei. Proc Natl Acad Sci U S A. 2001;98(15):8572-7.
- ↑ Pfeiffer V, Lingner J. TERRA promotes telomere shortening through exonuclease 1-mediated resection of chromosome ends. PLoS Genet. 2012;8(6):e1002747.
- 1 2 Luke B, Lingner J. TERRA: telomeric repeat-containing RNA. EMBO J 2009; 28:2503-10; PMID 19629047; http://dx.doi.org/10.1038/emboj.2009.166.
- ↑ Chawla R, Redon S, Raftopoulou C, Wischnewski H, Gagos S, Azzalin CM. Human UPF1 interacts with TPP1 and telomerase and sustains telomere leading-strand replication. EMBO J 2011; 30:4047- 58; PMID 21829167; http://dx.doi.org/10.1038/ emboj.2011.280.
- ↑ Feuerhahn S, Iglesias N, Panza A, Porro A, Lingner J. TERRA biogenesis, turnover and implications for func- tion. FEBS Lett 2010; 584:3812-8; PMID 20655916; http://dx.doi.org/10.1016/j.febslet.2010.07.032.
- ↑ Poveda AM, Le Clech M, Pasero P. Transcription and replication: breaking the rules of the road causes genomic instability. Transcription 2010; 1:99-102; PMID 21326900; http://dx.doi.org/10.4161/ trns.1.2.12665.
- ↑ Gómez-González B, García-Rubio M, Bermejo R, Gaillard H, Shirahige K, Marín A, et al. Genome- wide function of THO/TREX in active genes pre- vents R-loop-dependent replication obstacles. EMBO J 2011; 30:3106-19; PMID 21701562; http://dx.doi. org/10.1038/emboj.2011.206.
- ↑ Aguilera A, Gómez-González B. Genome instability: a mechanistic view of its causes and consequences. Nat Rev Genet 2008; 9:204-17; PMID 18227811; http:// dx.doi.org/10.1038/nrg2268.
- ↑ Flynn RL, Centore RC, O’Sullivan RJ, Rai R, Tse A, Songyang Z, Chang S, Karlseder J, Zou L. TERRA and hnRNPA1 orchestrate an RPA-to-POT1 switch on telomeric single-stranded DNA. Nature 2011, 471:532–536.
- 1 2 3 Ng L. J., Cropley J. E., Pickett H. A., Reddel R. R., Suter C. M. (2009). Telomerase activity is associated with an increase in DNA methylation at the proximal subtelomere and a reduction in telomeric transcription. Nucleic Acids Res. 37 1152–1159
- ↑ Chawla R, Azzalin C, M: The telomeric transcriptome and SMG proteins at the crossroads. Cytogenet Genome Res 2008;122:194-201. doi:10.1159/000167804
- ↑ de Silanes IL, d’Alcontres MS, Blasco MA. TERRA transcripts are bound by a complex array of RNA- binding proteins. Nat Commun 2010, 1:33.
- ↑ Artandi SE, DePinho RA. Telomeres and telom- erase in cancer. Carcinogenesis 2010; 31:9-18; PMID 19887512; http://dx.doi.org/10.1093/carcin/ bgp268.
- ↑ Blackburn EH. Walking the walk from genes through telomere maintenance to cancer risk. Cancer Prev Res (Phila) 2011; 4:473-5; PMID 21460394; http:// dx.doi.org/10.1158/1940-6207.CAPR-11-0066.
- ↑ Walne AJ, Dokal I. DyskeratosisCongenita: a his- torical perspective. Mech Ageing Dev 2008; 129:48- 59; PMID 18054794; http://dx.doi.org/10.1016/j. mad.2007.10.006.
- ↑ O’Sullivan RJ, Karlseder J. Telomeres: protecting chro- mosomes against genome instability. Nat Rev Mol Cell Biol 2010; 11:171-81; PMID 20125188.
- ↑ Armanios M. Syndromes of telomere shortening. Annu Rev Genomics Hum Genet 2009; 10:45-61; PMID 19405848; http://dx.doi.org/10.1146/annurev- genom-082908-150046.
- 1 2 Sampl S, Pramhas S, Stern C, Preusser M, Marosi C, Holzmann K. Expression of telomeres in astrocytoma WHO grade 2 to 4: TERRA level correlates with telomere length, telomerase activity, and advanced clinical grade. Transl On- col. 2012; 5: 56-65.
- ↑ Armanios M. Telomerase and idiopathic pulmonary fibrosis. Mutat Res 2012; 730:52-8; PMID 22079513; http://dx.doi.org/10.1016/j.mrfmmm.2011.10.013.
- ↑ Ehrlich M. The ICF syndrome, a DNA methyltransfer- ase 3B deficiency and immunodeficiency disease. Clin Immunol 2003; 109:17-28; PMID 14585272; http:// dx.doi.org/10.1016/S1521-6616(03)00201-8.
- 1 2 Deng Z, Campbell AE, Lieberman PM. TERRA, CpG methylation, and telomere heterochromatin: lessons from ICF syndrome cells. Cell Cycle 2010, 9:69–74.
- ↑ Nergadze SG, Farnung BO, Wischnewski H, Khoriauli L, Vitelli V, Chawla R, Giulotto E, Azzalin CM. CpG-island promoters drive transcription of human telomeres. RNA 2009, 15:2186–2194.
- ↑ Alcaín FJ, Villalba JM. Sirtuin activators. Expert Opin Ther Pat 2009; 19:403-14; PMID 19441923; http:// dx.doi.org/10.1517/13543770902762893.