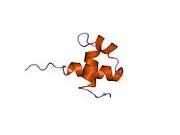
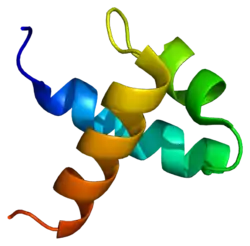
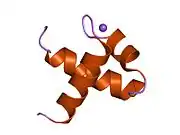
| Myb-like DNA-binding domain | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Identifiers | |||||||||
| Symbol | Myb_DNA-binding | ||||||||
| Pfam | PF00249 | ||||||||
| InterPro | IPR001005 | ||||||||
| PROSITE | PDOC00037 | ||||||||
| CATH | 1irz | ||||||||
| SCOP2 | 1irz / SCOPe / SUPFAM | ||||||||
| CDD | cd00167 | ||||||||
| |||||||||
Myb genes are part of a large gene family of transcription factors found in animals and plants. In humans, it includes Myb proto-oncogene like 1 and Myb-related protein B in addition to MYB proper.[5][6] Members of the extended SANT/Myb family also include the SANT domain and other similar all-helical homeobox-like domains.
Function
Viral
The Myb gene family is named after the eponymous gene in Avian myeloblastosis virus. The viral Myb (v-Myb, P01104) recognizes the sequence 5'-YAACKG-3'. It causes myeloblastosis (myeloid leukemia) in chickens.[7] Compared to the normal animal cellular Myb (c-myb), v-myb contains deletions in the C-terminal regulatory domain, leading to aberrant activation of other oncogenes.[8]
Animals
Myb proto-oncogene protein is a member of the MYB (myeloblastosis) family of transcription factors. The protein contains three domains, an N-terminal DNA-binding domain, a central transcriptional activation domain and a C-terminal domain involved in transcriptional repression. It may play a role in cell cycle regulation. Like the viral version, this gene is an oncogene, and rearrangements of the gene (often involving deletion in the C-terminal domain) causes cancer.[8]
Plants
MYB factors represent a family of proteins that include the conserved MYB DNA-binding domain. Plants contain a MYB-protein subfamily that is characterised by the R2R3-type MYB domain.[9]
In maize, phlobaphenes are synthesized in the flavonoids synthetic pathway[10] from polymerisation of flavan-4-ols[11][12] which encodes an R2R3 myb-like transcriptional activator[13] of the A1 gene encoding for the dihydroflavonol 4-reductase (reducing dihydroflavonols into flavan-4-ols)[14] while another gene (Suppressor of Pericarp Pigmentation 1 or SPP1) acts as a suppressor.[15] The maize P gene encodes a Myb homolog that recognizes the sequence CCWACC, in sharp contrast with the YAACGG bound by vertebrate Myb proteins.[16]
In sorghum, the corresponding yellow seed 1 gene (y1)[17] also encodes a R2R3 type of Myb domain protein that regulates the expression of chalcone synthase, chalcone isomerase and dihydroflavonol reductase genes required for the biosynthesis of 3-deoxyflavonoids.[18]
Ruby is a MYB transcriptional activator of genes that produce anthocyanin in citrus fruits. In most citrus varieties Ruby is non-functional, but in blood oranges it upregulates anthocyanin production to produce the characteristic red color of the fruit.[19]
See also
References
- 1 2 3 GRCh38: Ensembl release 89: ENSG00000118513 - Ensembl, May 2017
- 1 2 3 GRCm38: Ensembl release 89: ENSMUSG00000019982 - Ensembl, May 2017
- ↑ "Human PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ↑ "Mouse PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ↑ Chen Y, Xu H, Liu J, Zhang C, Leutz A, Mo X (Jul 2007). "The c-Myb functions as a downstream target of PDGF-mediated survival signal in vascular smooth muscle cells". Biochem Biophys Res Commun. 360 (2): 433–6. doi:10.1016/j.bbrc.2007.06.078. PMID 17599807.
- ↑ "Entrez Gene: v-myb myeloblastosis viral oncogene homolog (avian)".
- ↑ Klempnauer KH, Symonds G, Evan GI, Bishop JM (June 1984). "Subcellular localization of proteins encoded by oncogenes of avian myeloblastosis virus and avian leukemia virus E26 and by chicken c-myb gene". Cell. 37 (2): 537–47. doi:10.1016/0092-8674(84)90384-2. PMID 6327074. S2CID 46196167.
- 1 2 George OL, Ness SA (October 2014). "Situational awareness: regulation of the myb transcription factor in differentiation, the cell cycle and oncogenesis". Cancers. 6 (4): 2049–71. doi:10.3390/cancers6042049. PMC 4276956. PMID 25279451.
- ↑ Stracke R, Werber M, Weisshaar B (October 2001). "The R2R3-MYB gene family in Arabidopsis thaliana". Curr. Opin. Plant Biol. 4 (5): 447–56. doi:10.1016/s1369-5266(00)00199-0. PMID 11597504.
- ↑ Himi E, Mares DJ, Yanagisawa A, Noda K (July 2002). "Effect of grain colour gene (R) on grain dormancy and sensitivity of the embryo to abscisic acid (ABA) in wheat". J. Exp. Bot. 53 (374): 1569–74. doi:10.1093/jxb/erf005. PMID 12096095.
- ↑ Winkel-Shirley B (June 2001). "Flavonoid biosynthesis. A colorful model for genetics, biochemistry, cell biology, and biotechnology". Plant Physiol. 126 (2): 485–93. doi:10.1104/pp.126.2.485. PMC 1540115. PMID 11402179.
- ↑ Chopra S, Cocciolone SM, Bushman S, Sangar V, McMullen MD, Peterson T (March 2003). "The maize unstable factor for orange1 is a dominant epigenetic modifier of a tissue specifically silent allele of pericarp color1". Genetics. 163 (3): 1135–46. doi:10.1093/genetics/163.3.1135. PMC 1462483. PMID 12663550.
- ↑ Structural And Transcriptional Analysis Of The Complex P1-wr Cluster In Maize. Wolfgang Goettel, Joachim Messing. Plant & Animal Genomes XVI Conference Archived 2012-02-18 at the Wayback Machine
- ↑ Dong X, Braun EL, Grotewold E (September 2001). "Functional conservation of plant secondary metabolic enzymes revealed by complementation of Arabidopsis flavonoid mutants with maize genes" (PDF). Plant Physiol. 127 (1): 46–57. doi:10.1104/pp.127.1.46. hdl:1811/48809. PMC 117961. PMID 11553733.
- ↑ Lee EA, Harper V (2002). "Suppressor of Pericarp Pigmentation 1 (SPP1), a novel gene involved in phlobaphene accumulation in maize (Zea mays L.) pericarps". Maydica. 47 (1): 51–58. INIST 13772300
- ↑ Grotewold E, Drummond BJ, Bowen B, Peterson T (1994). "The myb-homologous P gene controls phlobaphene pigmentation in maize floral organs by directly activating a flavonoid biosynthetic gene subset". Cell. 76 (3): 543–554. doi:10.1016/0092-8674(94)90117-1. PMID 8313474. S2CID 42197232.
- ↑ Boddu J, Svabek C, Ibraheem F, Jones AD, Chopra S (2005). "Characterization of a deletion allele of a sorghum Myb gene yellow seedl showing loss of 3-deoxyflavonoids". Plant Science. 169 (3): 542–552. doi:10.1016/j.plantsci.2005.05.007. INIST 16983977
- ↑ Boddu J, Jiang C, Sangar V, Olson T, Peterson T, Chopra S (January 2006). "Comparative structural and functional characterization of sorghum and maize duplications containing orthologous myb transcription regulators of 3-deoxyflavonoid biosynthesis". Plant Mol. Biol. 60 (2): 185–99. doi:10.1007/s11103-005-3568-1. PMID 16429259. S2CID 23841582.
- ↑ Butelli E, Licciardello C, Zhang Y, Liu J, Mackay S, Bailey P, Reforgiato-Recupero G, Martin C (2012). "Retrotransposons control fruit-specific, cold-dependent accumulation of anthocyanins in blood oranges". Plant Cell. 24 (3): 1242–55. doi:10.1105/tpc.111.095232. PMC 3336134. PMID 22427337.
Further reading
- Oh IH, Reddy EP (1999). "The myb gene family in cell growth, differentiation and apoptosis". Oncogene. 18 (19): 3017–33. doi:10.1038/sj.onc.1202839. PMID 10378697.
- Nicolaides NC, Gualdi R, Casadevall C, Manzella L, Calabretta B (1991). "Positive autoregulation of c-myb expression via Myb binding sites in the 5' flanking region of the human c-myb gene". Mol. Cell. Biol. 11 (12): 6166–76. doi:10.1128/mcb.11.12.6166-6176.1991. PMC 361795. PMID 1944282.
- Kalkbrenner F, Guehmann S, Moelling K (1990). "Transcriptional activation by human c-myb and v-myb genes". Oncogene. 5 (5): 657–61. PMID 2189102.
- Westin EH, Gorse KM, Clarke MF (1990). "Alternative splicing of the human c-myb gene". Oncogene. 5 (8): 1117–24. PMID 2202948.
- Dasgupta P, Reddy EP (1990). "Identification of alternatively spliced transcripts for human c-myb: molecular cloning and sequence analysis of human c-myb exon 9A sequences". Oncogene. 4 (12): 1419–23. PMID 2687764.
- Janssen JW, Vernole P, de Boer PA, Oosterhuis JW, Collard JG (1986). "Sublocalization of c-myb to 6q21----q23 by in situ hybridization and c-myb expression in a human teratocarcinoma with 6q rearrangements". Cytogenet. Cell Genet. 41 (3): 129–35. doi:10.1159/000132217. PMID 3007038.
- Slamon DJ, Boone TC, Murdock DC, Keith DE, Press MF, Larson RA, Souza LM (1986). "Studies of the human c-myb gene and its product in human acute leukemias". Science. 233 (4761): 347–51. Bibcode:1986Sci...233..347S. doi:10.1126/science.3014652. PMID 3014652.
- Majello B, Kenyon LC, Dalla-Favera R (1987). "Human c-myb protooncogene: nucleotide sequence of cDNA and organization of the genomic locus". Proc. Natl. Acad. Sci. U.S.A. 83 (24): 9636–40. doi:10.1073/pnas.83.24.9636. PMC 387195. PMID 3540945.
- Szczylik C, Skorski T, Ku DH, Nicolaides NC, Wen SC, Rudnicka L, Bonati A, Malaguarnera L, Calabretta B (1993). "Regulation of proliferation and cytokine expression of bone marrow fibroblasts: role of c-myb". J. Exp. Med. 178 (3): 997–1005. doi:10.1084/jem.178.3.997. PMC 2191153. PMID 7688794.
- Krieg J, Oelgeschläger M, Janknecht R, Lüscher B (1995). "High affinity DNA binding of native full length c-Myb and differential proteolytic sensitivity of its N- and C-terminal domains". Oncogene. 10 (11): 2221–8. PMID 7784067.
- Jacobs SM, Gorse KM, Westin EH (1994). "Identification of a second promoter in the human c-myb proto-oncogene". Oncogene. 9 (1): 227–35. PMID 8302584.
- Favier D, Gonda TJ (1994). "Detection of proteins that bind to the leucine zipper motif of c-Myb". Oncogene. 9 (1): 305–11. PMID 8302594.
- Glaser R, Lafuse WP, Bonneau RH, Atkinson C, Kiecolt-Glaser JK (1993). "Stress-associated modulation of proto-oncogene expression in human peripheral blood leukocytes". Behav. Neurosci. 107 (3): 525–9. doi:10.1037/0735-7044.107.3.525. PMID 8329139.
- Dash AB, Orrico FC, Ness SA (1996). "The EVES motif mediates both intermolecular and intramolecular regulation of c-Myb". Genes Dev. 10 (15): 1858–69. doi:10.1101/gad.10.15.1858. PMID 8756344.
- Vorbrueggen G, Lovrić J, Moelling K (1997). "Functional analysis of phosphorylation at serine 532 of human c-Myb by MAP kinase". Biol. Chem. 377 (11): 721–30. doi:10.1515/bchm3.1996.377.11.721. PMID 8960373.
- Tavner FJ, Simpson R, Tashiro S, Favier D, Jenkins NA, Gilbert DJ, Copeland NG, Macmillan EM, Lutwyche J, Keough RA, Ishii S, Gonda TJ (1998). "Molecular cloning reveals that the p160 Myb-binding protein is a novel, predominantly nucleolar protein which may play a role in transactivation by Myb". Mol. Cell. Biol. 18 (2): 989–1002. doi:10.1128/MCB.18.2.989. PMC 108811. PMID 9447996.
- Hedge SP, Kumar A, Kurschner C, Shapiro LH (1998). "c-Maf interacts with c-Myb to regulate transcription of an early myeloid gene during differentiation". Mol. Cell. Biol. 18 (5): 2729–37. doi:10.1128/mcb.18.5.2729. PMC 110652. PMID 9566892.
- Leverson JD, Koskinen PJ, Orrico FC, Rainio EM, Jalkanen KJ, Dash AB, Eisenman RN, Ness SA (1998). "Pim-1 kinase and p100 cooperate to enhance c-Myb activity". Mol. Cell. 2 (4): 417–25. doi:10.1016/S1097-2765(00)80141-0. PMID 9809063.
- Thompson MA, Rosenthal MA, Ellis SL, Friend AJ, Zorbas MI, Whitehead RH, Ramsay RG (1998). "c-Myb down-regulation is associated with human colon cell differentiation, apoptosis, and decreased Bcl-2 expression". Cancer Res. 58 (22): 5168–75. PMID 9823328.
- Verbeek W, Gombart AF, Chumakov AM, Müller C, Friedman AD, Koeffler HP (1999). "C/EBPepsilon directly interacts with the DNA binding domain of c-myb and cooperatively activates transcription of myeloid promoters". Blood. 93 (10): 3327–37. doi:10.1182/blood.V93.10.3327. PMID 10233885.
External links
- MYB+protein,+human at the U.S. National Library of Medicine Medical Subject Headings (MeSH)
- Drosophila Myb oncogene-like - The Interactive Fly
- Arabidospsis thaliana MYB family at Database of Arabidopsis Transcription Factors (DATF)
This article incorporates text from the United States National Library of Medicine, which is in the public domain.